- The paper introduces DIM-GCN, which disentangles disease-relevant and irrelevant factors via a dual-encoder architecture and graph convolutional networks.
- It employs Bayesian variational inference and structured clinical priors to enhance out-of-distribution robustness in mammogram classification.
- Empirical results demonstrate up to a 6.2% AUC improvement over ERM across diverse hospital datasets, highlighting its practical efficacy.
Domain-Invariant Mammogram Classification via Graph Convolutional Networks
Introduction
Cross-domain robustness is critical for automatic mammogram classification due to the notorious domain shifts induced by varying imaging conditions and clinical protocols. Classical empirical risk minimization (ERM) fails under such distribution shifts, as it often entangles disease-irrelevant biases (e.g., hospital-specific post-processing artifacts) with disease-relevant features, which degrades generalization on out-of-distribution (OOD) cases.
This work introduces a Domain Invariant Model with Graph Convolutional Network (DIM-GCN) for robust mammogram patch-level benign/malignant mass classification. DIM-GCN explicitly separates disease-relevant and -irrelevant latent factors and harnesses attribute-level medical priors using GCNs for improved identifiability and transfer.
Macroscopic and Microscopic Priors
Clinical evidence dictates that both macroscopic (shape, margin, spiculation) and microscopic (texture, curvature) features determine malignancy. These visual attributes, summarized in standards such as BI-RADS, provide domain-invariant signals, while irrelevant confounds—stemming from, e.g., acquisition hardware—are hospital-specific.
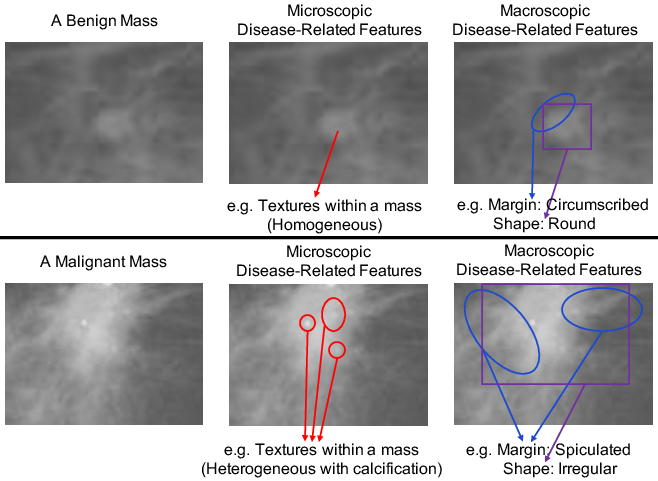
Figure 1: Macroscopic and microscopic indicators in mammogram patches: benign masses are homogeneous, circumscribed, and regular; malignant ones are heterogeneous, spiculated, and irregular.
DIM-GCN's architecture encodes these priors, separately modeling macroscopic and microscopic features, and imposes explicit invariance through the training objective.
Bayesian Decomposition and Disentanglement
DIM-GCN leverages a Bayesian network to decompose latent factors generating images into three groups: a (macroscopic), s (microscopic), and z (disease-irrelevant, domain-dependent). Only a and s causally depend on the disease label; z encodes imaging biases and is domain-specific.
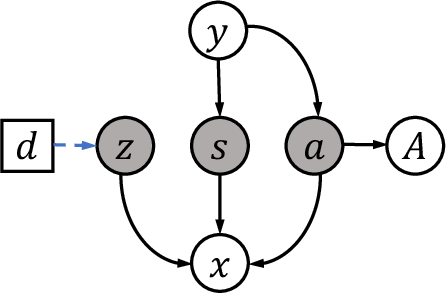
Figure 2: Causal graph: latent factors a (macroscopic), s (microscopic), and z (domain-irrelevant) jointly generate images; only a, s are linked to the disease label and attributes.
A formal identifiability theorem ensures that, given sufficient domain diversity, disease-relevant representations (a,s) can be universally separated from z in the generative mechanism, up to invertible transformations. This theoretical result underpins the entire method.
Variational Inference and Network Design
DIM-GCN employs a domain-conditional variational autoencoder. The encoder comprises two branches: a domain-invariant path extracting a,s (the relevant encoder), and a domain-specific path for z (the irrelevant encoder), with domain-adaptive normalization to handle multiple domains. At test time, only the relevant encoder is used, guaranteeing invariance to unseen domains.
To best leverage macroscopic priors, a is regularized to reconstruct clinical attributes through a GCN that captures attribute interdependence, reflecting structured clinical knowledge.
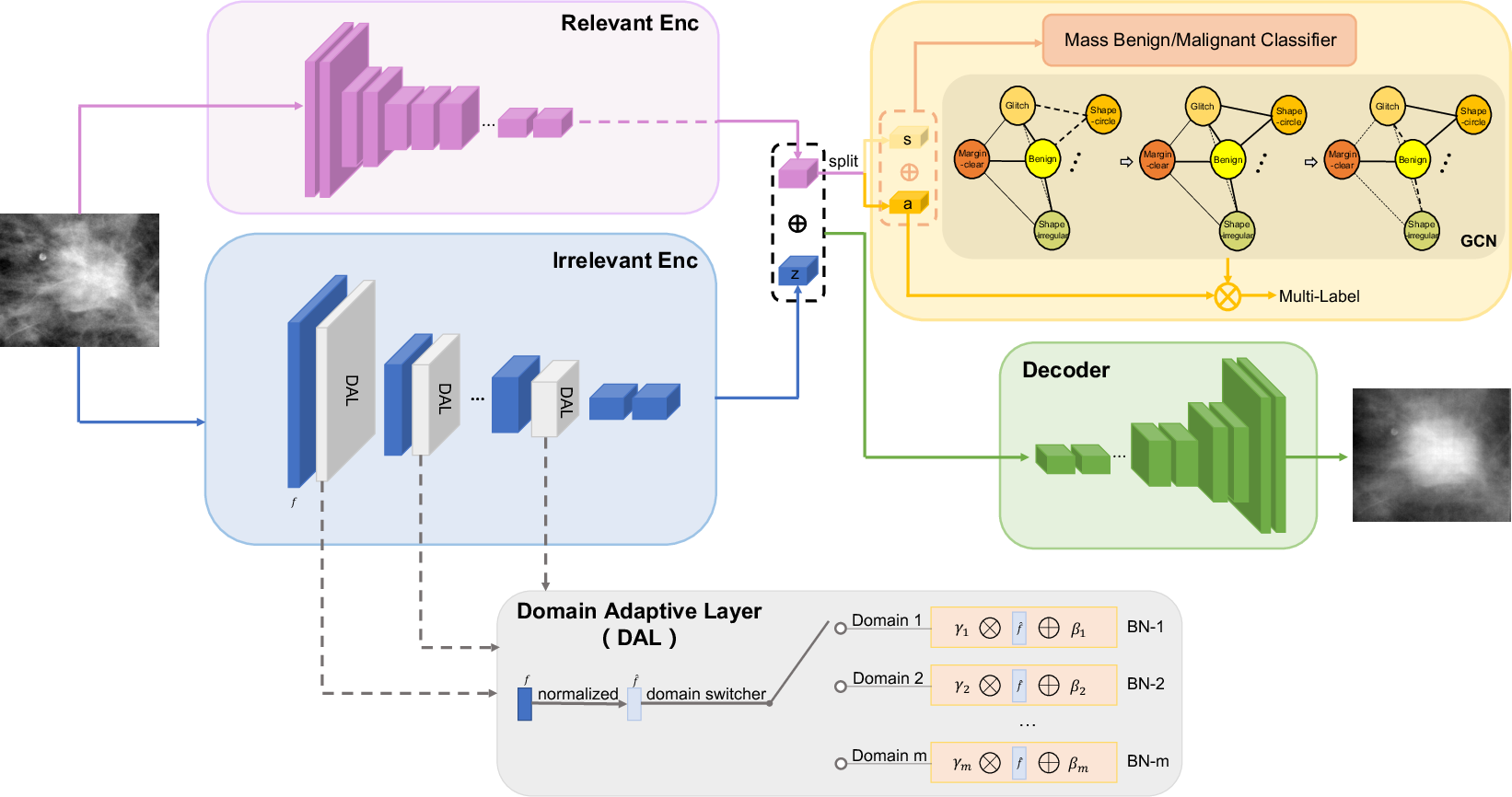
Figure 3: DIM-GCN architecture: training leverages both relevant and domain-adaptive irrelevant encoders; GCN reconstructs clinical attributes from a. Inference on unseen domains uses only the relevant encoder.
Reconstruction losses on x, attribute losses on A (via GCN), and classification losses on y (using a,s only) together, accompanied by a variance regularizer enforcing statistical invariance of a,s across domains, constitute the compound objective.
Experimental Protocol
Evaluation included the public DDSM dataset and three in-house mammogram datasets, each representing a distinct hospital/domain. OOD settings were enforced by training on three domains and testing on the fourth. The primary metric was AUC for benign/malignant classification.
Numerous baselines were compared: ERM, adversarial training (DANN, MMD-AAE), disentanglement (Guided-VAE, DIVA), attribute-guided models (Chen et al., ICADx), and invariant risk minimization.
Results and Analysis
DIM-GCN demonstrates consistent superiority across all OOD tasks, improving average AUC by up to 6.2% over ERM and finishing ahead of all listed baselines. Notably, the method approaches the in-domain AUC upper bounds, while standard methods suffer significant OOD degradation.
Ablations confirm the criticality of (i) both relevant and irrelevant encoder branches, (ii) structured GCN-based attribute supervision, (iii) explicit disentanglement of s, and (iv) variance regularization. Reducing the ratio of domain-adaptive layers or removing explicit disentanglement leads to pronounced performance drops.
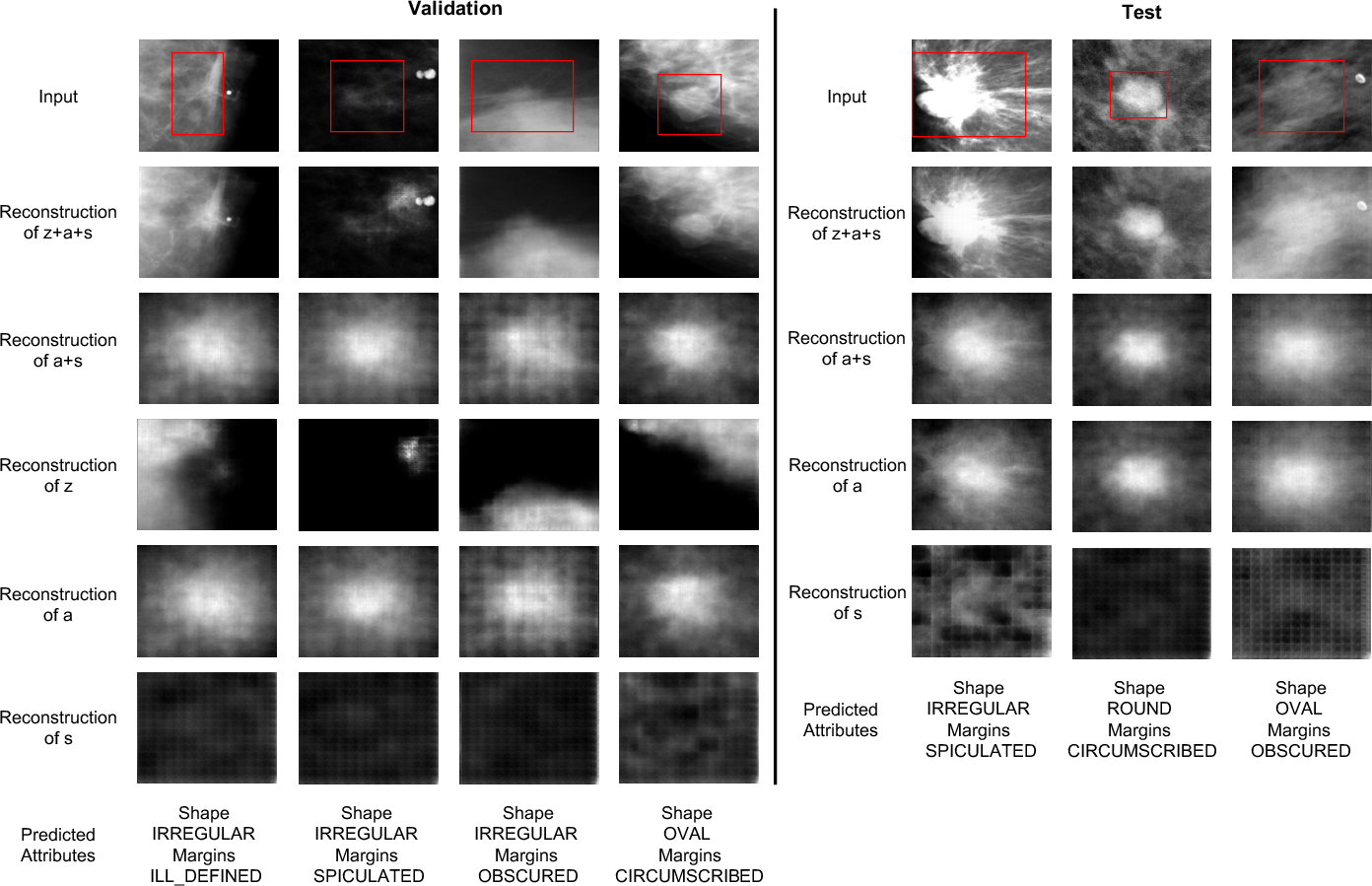

Figure 4: Reconstructions under OOD setting. Lesion-related features (a,s) faithfully reconstruct the diseased area, while disease-irrelevant z captures nuisance information such as background and acquisition artifacts.
Further, DIM-GCN excels at attribute prediction (multi-label classification of shape and margin), indicating effective capture of clinical priors and interpretable feature extraction that aligns with radiological practice.
Theoretical and Practical Implications
DIM-GCN's disentanglement theorem provides a rigorous basis for learning transferable, causal representations in high-stakes medical imaging. The split encoder architecture and attribute-constrained GCN offer practical value: the approach ablates domain-specific biases, clinical factors are explicitly modeled, and the model is robust under cross-hospital deployment. This paradigm is extensible to other medical imaging domains plagued by spurious correlations and confounding.
Future Directions
Potential developments include extending causal disentanglement frameworks to other diagnostic modalities (e.g., CT, MRI), integrating richer clinical knowledge graphs for attribute supervision, and scaling to weakly supervised or semi-supervised OOD generalization scenarios.
Conclusion
DIM-GCN establishes a novel paradigm for robust mammogram diagnosis, combining causal invariance, clinical attribute modeling via GCNs, and disentangled variational inference. Its empirical and theoretical contributions set the groundwork for more principled, deployable AI systems in medical image analysis, where domain shifts are an inevitably present challenge.



