SCP: Accelerating Discovery with a Global Web of Autonomous Scientific Agents
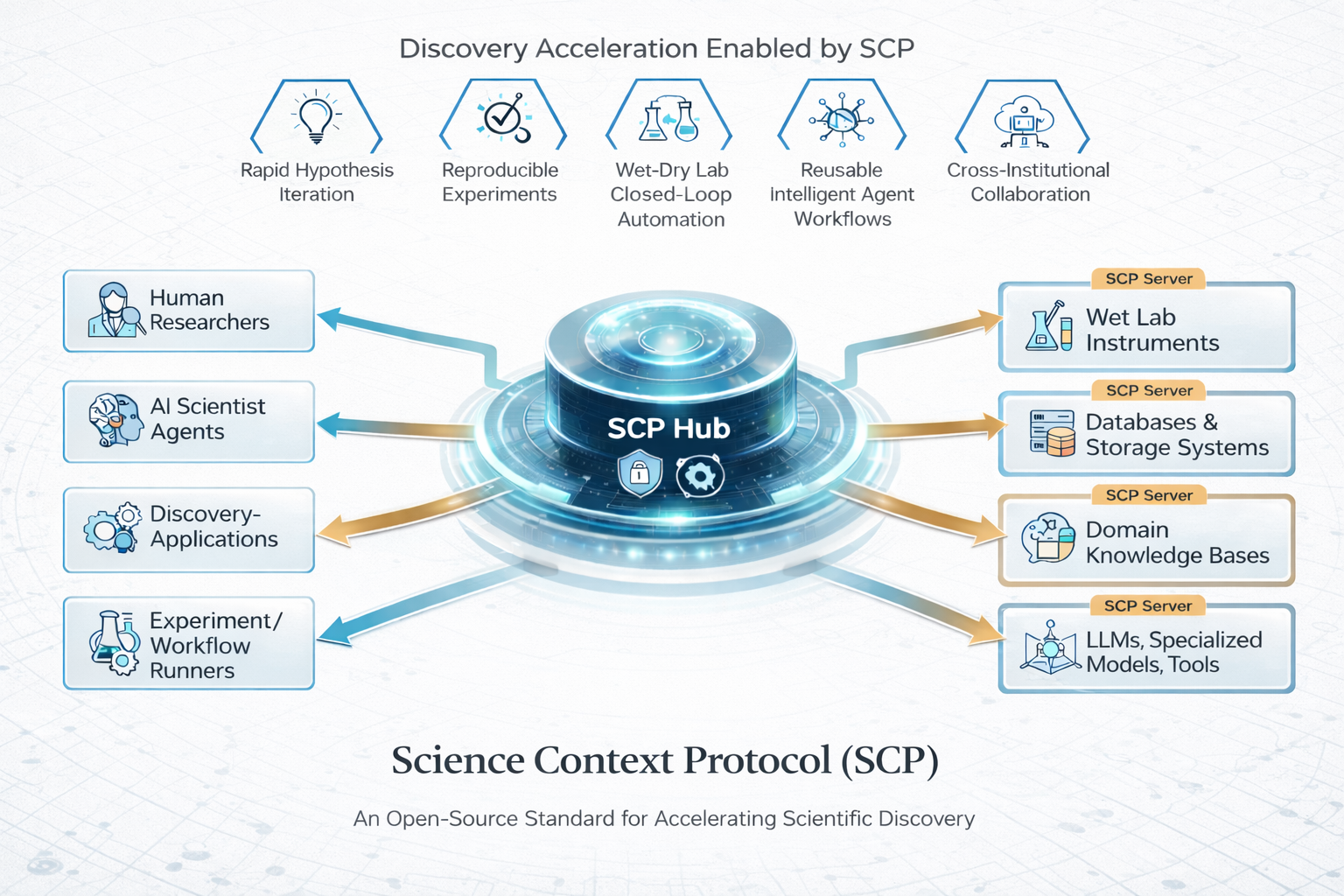
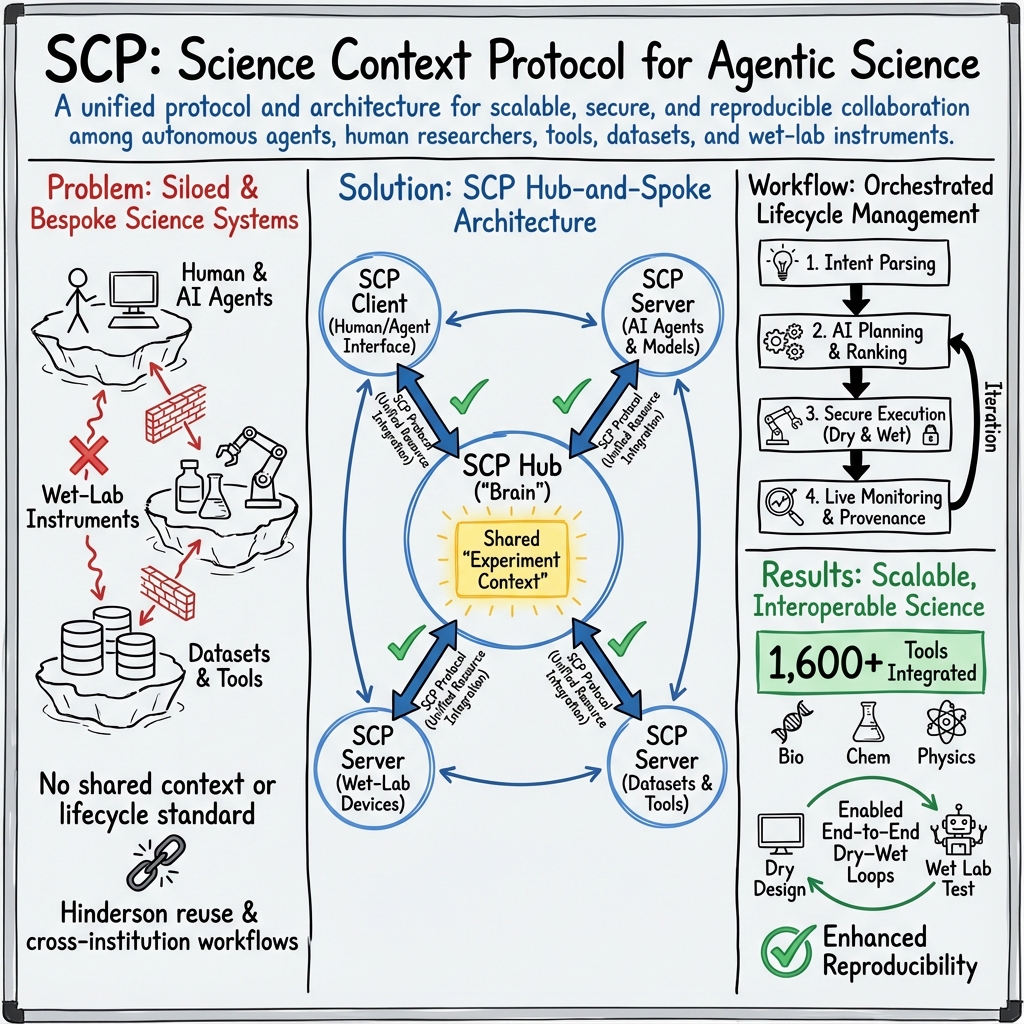
Abstract: We introduce SCP: the Science Context Protocol, an open-source standard designed to accelerate discovery by enabling a global network of autonomous scientific agents. SCP is built on two foundational pillars: (1) Unified Resource Integration: At its core, SCP provides a universal specification for describing and invoking scientific resources, spanning software tools, models, datasets, and physical instruments. This protocol-level standardization enables AI agents and applications to discover, call, and compose capabilities seamlessly across disparate platforms and institutional boundaries. (2) Orchestrated Experiment Lifecycle Management: SCP complements the protocol with a secure service architecture, which comprises a centralized SCP Hub and federated SCP Servers. This architecture manages the complete experiment lifecycle (registration, planning, execution, monitoring, and archival), enforces fine-grained authentication and authorization, and orchestrates traceable, end-to-end workflows that bridge computational and physical laboratories. Based on SCP, we have constructed a scientific discovery platform that offers researchers and agents a large-scale ecosystem of more than 1,600 tool resources. Across diverse use cases, SCP facilitates secure, large-scale collaboration between heterogeneous AI systems and human researchers while significantly reducing integration overhead and enhancing reproducibility. By standardizing scientific context and tool orchestration at the protocol level, SCP establishes essential infrastructure for scalable, multi-institution, agent-driven science.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
What is this paper about?
This paper introduces SCP, the Science Context Protocol. Think of SCP as a “universal language and traffic controller” that helps AI systems and people work together with scientific tools, data, and real lab machines. Its goal is to make science faster and more reliable by letting different labs and AI agents easily connect, plan, run, and track experiments in a consistent and safe way—like turning many separate labs and apps into one big, cooperative web.
What questions does it try to answer?
The paper focuses on a few simple questions:
- How can we get AI “scientists” and human researchers to share tools, data, and lab equipment smoothly across different places?
- How can we describe any scientific tool (a software model, a database, or even a robot in a lab) in one standard way so it’s easy to find and use?
- How can we plan and run whole experiments—from idea to results—safely, with good records, and with less manual setup?
- How can we make this work not just for computer-only (“dry”) work, but also for real-world lab (“wet”) experiments?
How did the researchers build and test it?
The team built SCP with two big ideas in mind:
- Unified Resource Integration: A standard way to describe and call any scientific resource. Imagine a universal plug that fits every tool—software, data, AI models, and lab devices—so agents know what each tool can do and how to use it.
- Orchestrated Experiment Lifecycle: A system that guides an experiment from start to finish—registering it, planning steps, executing tasks, monitoring progress, handling errors, and saving everything for later reuse.
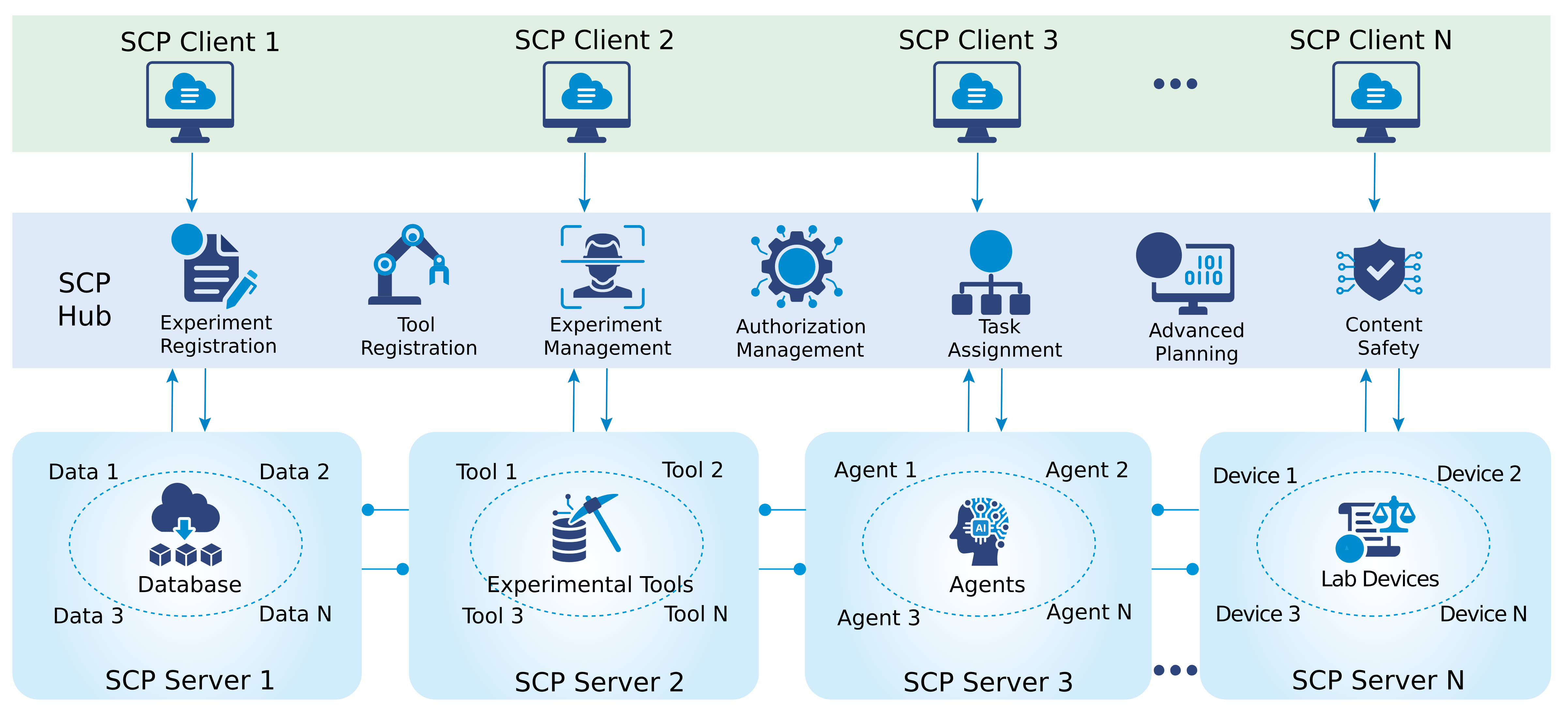
To make this work in practice, SCP uses a “hub-and-spoke” design:
- SCP Hub (the “brain”): Plans experiments from high-level goals, finds the right tools, schedules tasks, and keeps detailed records. It also handles security and permissions so only the right people and agents can do certain actions.
- SCP Servers (the “hands”): These run near tools and lab devices. They receive tasks from the Hub and carry them out—calling an AI model, running a simulation, moving a lab robot, or reading a sensor—and send results back.
- SCP Clients (the “remote control”): Interfaces that people or AI agents use to ask for experiments, review plans, watch progress live, and see results.
Key features explained in everyday language:
- Protocol: A shared set of rules so everything speaks the same language.
- Orchestration: Like a conductor guiding an orchestra—deciding who plays when, checking if the music is right, and fixing issues mid-performance.
- Experiment context: A “project folder” that stores what the goal is, what steps were planned, what actually happened, and all results, so experiments are traceable and repeatable.
- Dry–wet integration: Treating computer tools and real lab machines as first-class citizens under the same system.
- Safety and security: Fine-grained permissions and checks so experiments are done responsibly and data stays protected.
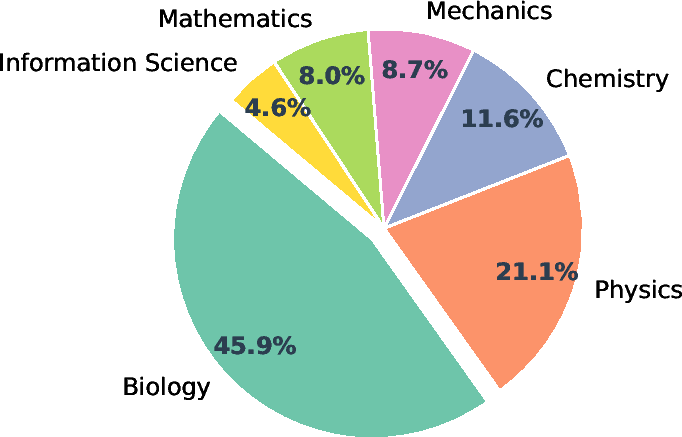
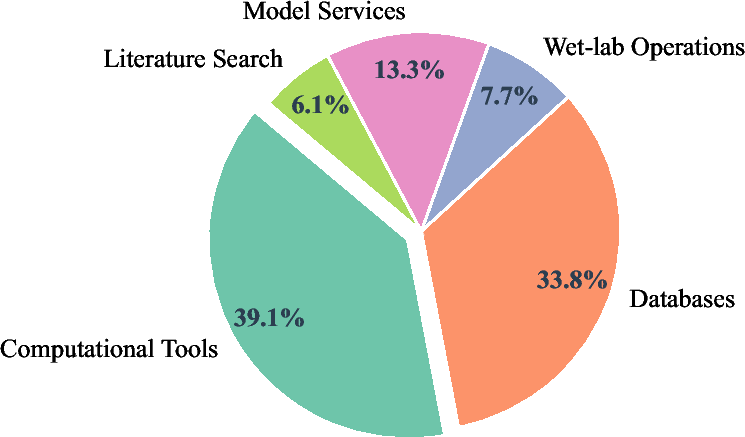
They also built a large open platform called Intern-Discovery that connects 1,600+ tools through SCP, covering areas like biology, chemistry, materials, physics, data processing, and machine learning.
What did they find, and why is it important?
Main results:
- A working open-source protocol and system that lets AI agents and humans discover, combine, and run tools across labs—computers and robots alike—under one consistent setup.
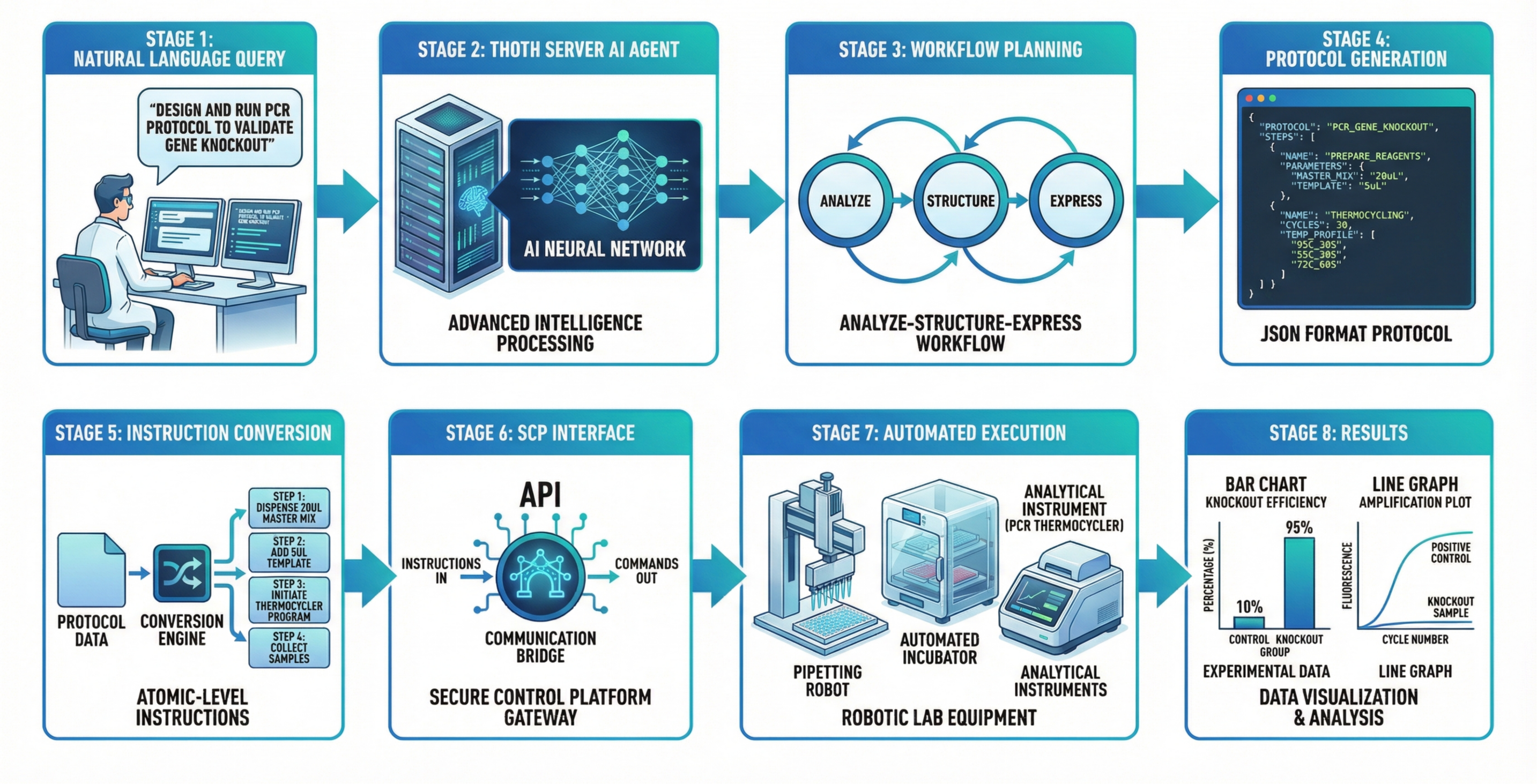
- An intelligent planner in the Hub that can turn a plain request (“run a PCR to check a gene knockout”) into a detailed, step-by-step plan, rank different ways to do it, estimate time and cost, and then execute it with monitoring and automatic error handling.
- A big ecosystem of 1,600+ tools connected through SCP, so agents can quickly build complex workflows without custom glue code for each new tool.
- Strong tracking and reproducibility: every action and result is recorded in a standard format, making it easier to repeat or audit experiments.
- Smooth collaboration across institutions: the Hub and Servers work over secure channels and respect local permissions, so multiple labs can safely share capabilities.
Why it matters:
- Less time wiring systems together; more time doing science.
- Fewer errors thanks to planning, validation, and monitoring.
- Reproducible experiments by default, which is key for trustworthy science.
- Easier teamwork between people, AI agents, and lab robots—even across different labs and tools.
The paper also shows several real examples:
- Automatic protocol design and execution: The system turns a high-level request into a machine-readable lab protocol and carries it out on instruments.
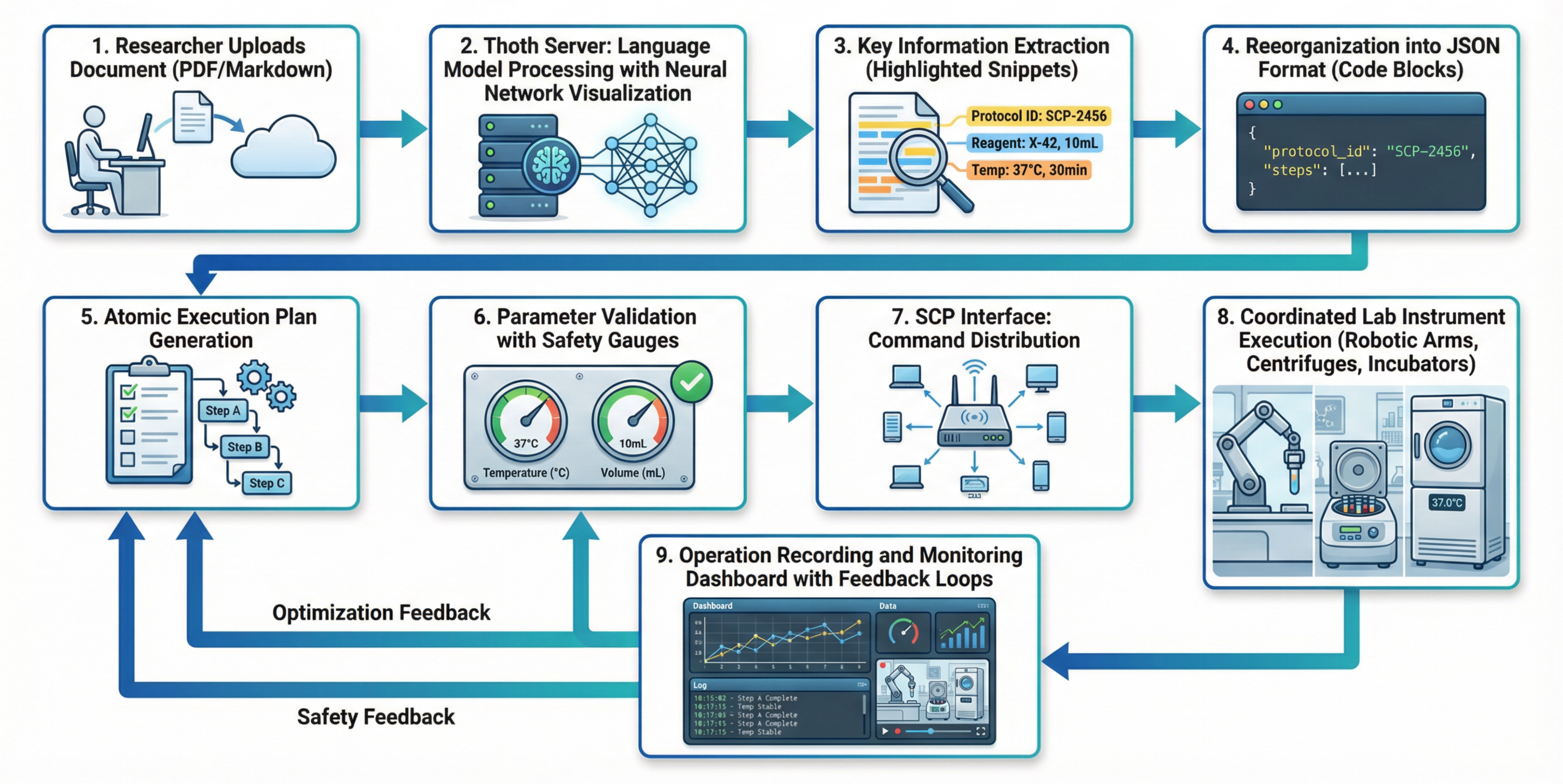
- Re-running a protocol from a PDF: Upload a methods document, and SCP converts it into an executable plan for lab robots—no manual rewriting.
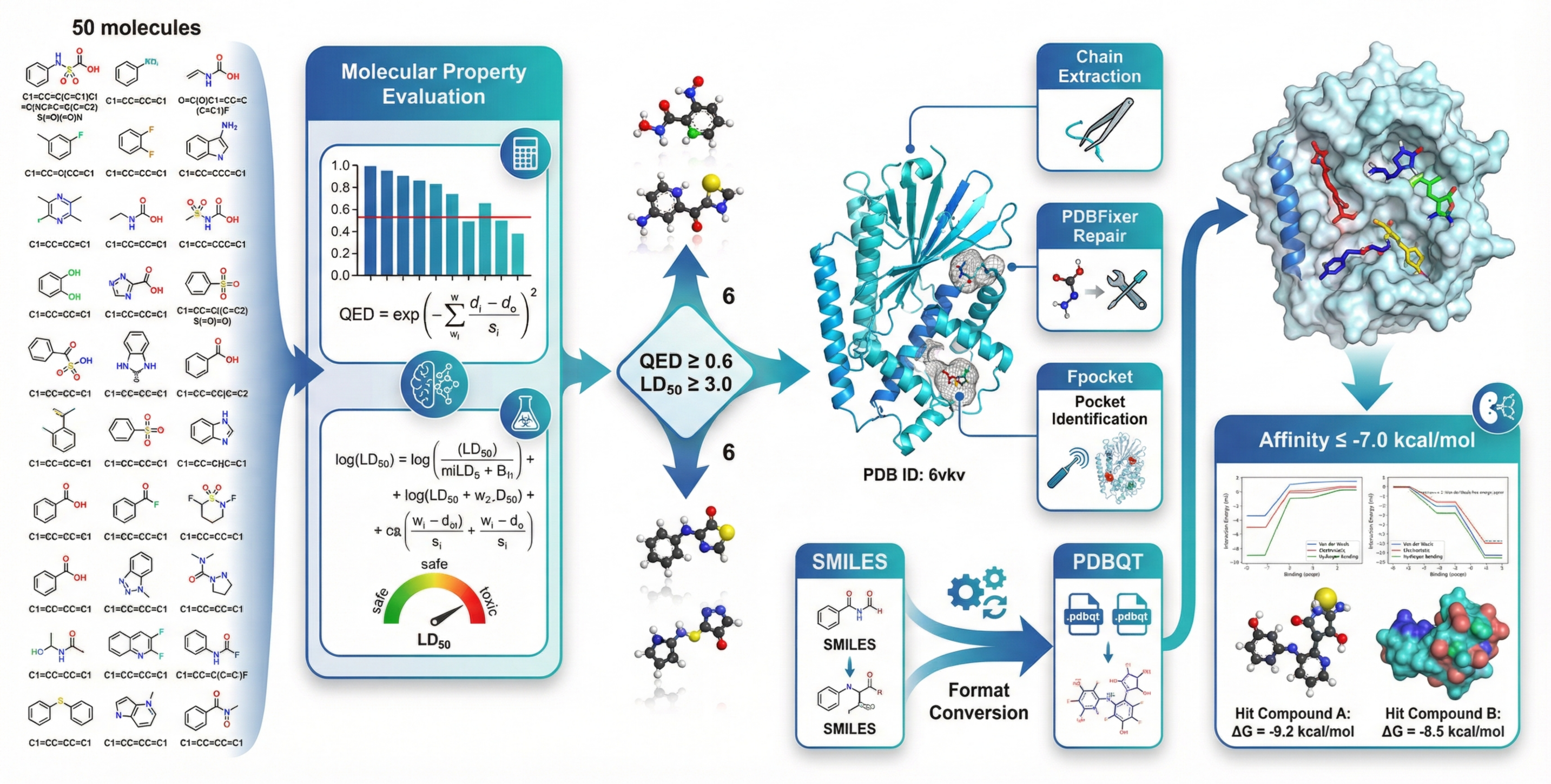
- AI-driven drug discovery steps: From filtering candidate molecules to docking them to a protein target, SCP runs the chain of tools end-to-end.
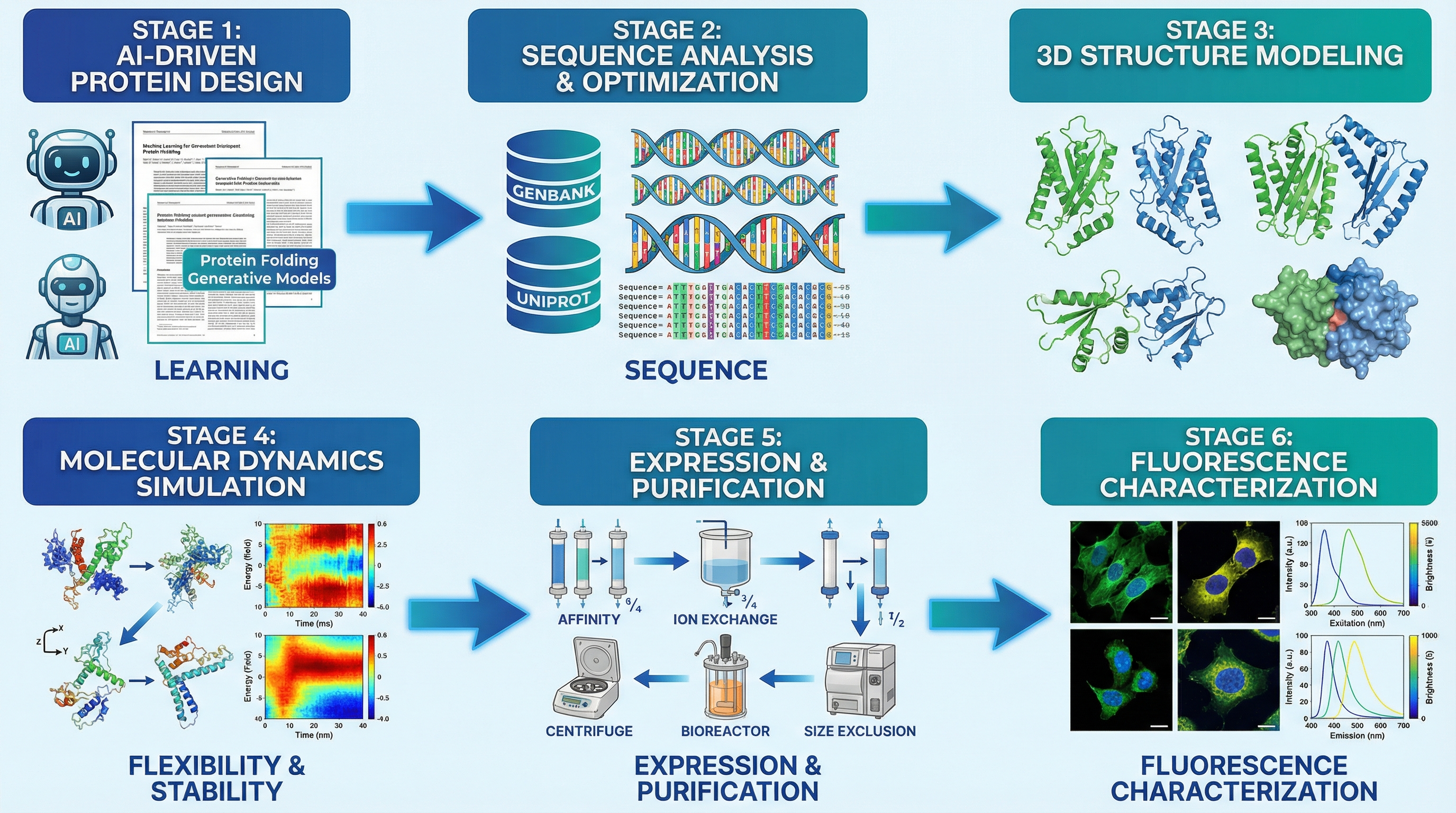
- Protein engineering loop: Designs variants on computers, tests them on lab robots, and feeds results back to improve the next round—all in one connected workflow.
What is the impact, and what could happen next?
SCP could become core infrastructure for “agent-driven science,” where smart software and lab automation help scientists test ideas faster and more safely. Possible impacts include:
- Faster discovery: Planning, running, and refining experiments becomes quicker and more reliable.
- Bigger collaboration: Labs can share tools and data through a common protocol while keeping control over who can do what.
- Better science quality: Clear records and repeatable workflows increase trust in results.
- Easier scaling: As more tools and devices adopt SCP, building new scientific workflows becomes like snapping together Lego bricks.
In short, SCP aims to do for scientific experimentation what standards like USB and the web did for computers—make everything connect, communicate, and work together, so innovation can move much faster.
Knowledge Gaps
Knowledge Gaps, Limitations, and Open Questions
Below is a single, concrete list of what remains missing, uncertain, or unexplored in the paper, aimed at guiding future research and development.
- Lack of quantitative system-level evaluation (e.g., end-to-end latency, throughput, failure rates, recovery time) for dry–wet workflows under realistic multi-institution loads.
- No benchmarking against existing orchestration frameworks (including MCP-based systems) on standardized tasks to substantiate claimed advantages.
- Unspecified algorithms and models for plan synthesis and ranking (top-k selection): planning formalism, cost/latency/risk estimation methods, calibration, and validation metrics.
- Absent formal protocol specification for experiment plans beyond “JSON schema”: no defined semantics for preconditions, postconditions, side effects, invariants, or effect typing.
- Missing verification and validation strategy for AI-generated protocols (e.g., static checks, model checking, simulation-in-the-loop, formal safety constraints) prior to wet-lab execution.
- Insufficient detail on anomaly detection: signal sources, thresholds, models used, false-positive/negative rates, and evaluation on real-world incident data.
- Rollback semantics for physical actions are unclear (what can be safely undone, how state is tracked, how partial rollbacks avoid compounding risks).
- Resource scheduling is under-specified (queuing, priority, fairness, preemption, multi-tenant isolation), and lacks algorithmic description and empirical evaluation.
- No clear resilience plan for network partitions, Server or Hub outages, or device unavailability (failover strategies, state reconciliation, disaster recovery).
- Centralized Hub poses a single point of failure/trust: missing design and evaluation for federation, multi-hub coordination, consensus, and cross-hub context synchronization.
- Sparse security architecture beyond OAuth2.1: lacking details on fine-grained authorization (RBAC/ABAC), secret management, multi-tenant isolation, and audit log integrity.
- No attestation or trust model for edge Servers and tools (e.g., code provenance, signed drivers, sandboxing, runtime isolation, supply-chain security for third-party adapters).
- Unexplored cryptographic provenance and tamper-evident audit trails (e.g., W3C PROV alignment, hash-chaining, secure timestamping) for regulatory-grade traceability.
- Data privacy and regulatory compliance not specified (GDPR, HIPAA, IRB/ethics, biosecurity/dual-use policies, cross-border data residency, export control).
- Wet-lab safety controls are not detailed (interlocks, hazard assessment, kill-switches, human-in-the-loop policies, device calibration verification, safety certifications).
- No alignment to established scientific ontologies and standards (FAIR principles, OBI, CHEBI, MIAME/OME, ISA-Tab) for interoperable context and metadata exchange.
- Versioning and evolution of the SCP schemas are not described (backward compatibility, migration paths, capability negotiation, deprecation policies).
- Interoperability with MCP and other protocols lacks concrete adapters/mappings, migration strategies, and empirical tests of cross-protocol workflows.
- Tool discovery at scale (1,600+ tools) lacks evaluation of search/ranking quality, metadata completeness, duplicate resolution, and cold-start integration overhead.
- Tool curation and lifecycle management are undefined (quality assurance, versioning, deprecation, licensing constraints, security review, continuous integration testing).
- Integration with ELN/LIMS, HPC schedulers (SLURM, PBS), and containerized environments (Docker/Singularity) is claimed conceptually but lacks concrete APIs and performance data.
- Absent quantitative results in case studies (e.g., task duration, error rates, yield improvements, docking accuracy vs. baselines) and independent replication.
- Explainability of orchestration decisions is not evaluated (usefulness, fidelity of rationales, human-understandability, effect on trust and oversight).
- Human factors and usability are unassessed (training needs, cognitive load, UI design, operator error rates, effectiveness of dashboards and alerts).
- Economic analysis is missing (cost models for compute/instrument time, budget forecasting accuracy, cost–benefit vs. traditional workflows).
- Concurrency control for multi-agent collaboration is unspecified (shared context consistency, conflict resolution, locking/version control, knowledge synchronization).
- Real-time constraints for device control are not addressed (timing guarantees, jitter, clock synchronization, determinism, integration with ICS/OT safety standards).
- Open questions around cross-institution governance (data-sharing agreements, IP management, attribution, incentive mechanisms, compliance auditing) remain unaddressed.
- No standard test suite or conformance program for SCP implementations (reference datasets, plugfests, fuzz testing, certification criteria).
- Unclear strategy for continuous learning from past experiments (experience replay, meta-learning for planning, bias detection, and mitigation).
- Risks from LLM hallucinations and agent misalignment are acknowledged implicitly but not formally mitigated (guardrails, red-teaming, domain-specific verification).
- Unknown scalability limits of the Hub registry and orchestration under high tool/device churn (registration storms, health-check scaling, routing bottlenecks).
- Edge execution environment details are missing (isolation of device drivers, resource quotas, privilege boundaries, safe API bridging to proprietary vendor controllers).
- Lack of community governance and standardization roadmap (working groups, change control, stewardship model, adoption incentives, alignment with standards bodies).
- Unexplored legal and ethical safeguards for dual-use concerns in chemistry and biology (automated synthesis protocols, access controls for hazardous operations).
- Unclear interoperability with data lakes and streaming systems (schema-on-read/write, lineage propagation, time-series handling, eventual consistency across logs).
- No documented performance or accuracy of risk predictions used in plan ranking (calibration error, domain generalization, stress testing).
- Unspecified monitoring SLOs/SLA (uptime targets, alerting thresholds, mean time to detection/repair, performance baselines for critical paths).
Practical Applications
Below are actionable, real-world applications that flow directly from the SCP protocol, architecture, and case studies described in the paper. They are grouped by immediacy and annotated with sectors, potential products/workflows, and key dependencies that may affect feasibility.
Immediate Applications
- Cross-lab reproducibility and provenance-by-default for experiments Sectors: academia, pharma/biotech, CROs, national labs Tools/products/workflows: SCP Hub integrated with ELN/LIMS; JSON experiment plans; per-experiment IDs; audit logs; “provenance export” for compliance packages Assumptions/dependencies: OAuth2.1/IAM integration; data-governance mappings; stable SCP schema; institutional buy-in for centralized Hub logging
- Automated protocol ingestion and replay from PDFs (Case Study 2) Sectors: biotech R&D, CROs, publishing/journals, teaching labs Tools/products/workflows: Thoth protocol understanding; Thoth-OP execution planning; SCP Hub dispatch to robots/instruments; “PDF→Executable Protocol” service Assumptions/dependencies: quality of protocol extraction on noisy PDFs; validated device drivers; safety interlocks; human-in-the-loop approval for high-risk steps
- End-to-end closed-loop wet-lab orchestration from high-level intent (Case Study 1) Sectors: synthetic biology, bioprocess, analytics labs, diagnostics development Tools/products/workflows: experiment-flow API; plan ranking and execution; anomaly detection/rollback; live dashboards; “Hub-as-a-brain” controlling connected SCP Servers Assumptions/dependencies: instrument APIs and calibration; facility safety SOPs; reliable network; fallback/rollback strategies configured
- AI-driven small-molecule triage and docking pipelines (Case Study 3) Sectors: pharma/biotech (hit identification), computational chemistry CROs Tools/products/workflows: QED/ADMET evaluators; fpocket; Vina-style docking via SCP; “SMILES-in → ranked hits-out” workflow templates Assumptions/dependencies: model quality and domain validation; compute/HPC access; dataset licensing; alignment with medicinal chemistry decision criteria
- Integrated dry–wet protein engineering loops (Case Study 4) Sectors: enzyme engineering, fluorescent protein optimization, synthetic biology tools Tools/products/workflows: sequence design + structure/property prediction + robotic assays in one SCP context; variant ranking; condition optimization loops Assumptions/dependencies: accurate in silico predictors for target properties; assay reproducibility; throughput and reliability of automation platforms
- Multi-institution tool and instrument discovery with fine-grained access control Sectors: research consortia, industry–academia partnerships Tools/products/workflows: SCP Hub registry; federated SCP Servers; per-resource ACLs; cross-org service discovery; “bring-your-own-server” onboarding Assumptions/dependencies: legal/data-sharing agreements; network security; standardized metadata; resource usage accounting
- “Tool galaxy” access for LLM agents (1,600+ tools) under a single schema Sectors: software for scientific agents, lab informatics, research IT Tools/products/workflows: tool registry and search; capability matchmaking; plug-and-play toolkits for bio/chem/materials; agent SDKs using SCP Assumptions/dependencies: ongoing curation; versioning and deprecation policy; schema conformance by tool maintainers
- HPC and cloud simulation orchestration in the same experiment context Sectors: materials science, climate/earth science, CFD/FEA in engineering Tools/products/workflows: job submission via SCP Servers; data staging and provenance; “simulation + wet validation” hybrid protocols Assumptions/dependencies: connectors to schedulers (SLURM, Kubernetes); data locality strategies; cost controls and quotas
- Compliance-ready audit, monitoring, and risk controls Sectors: regulated labs (GxP), diagnostics, medical devices Tools/products/workflows: immutable experiment logs; permissioned data stores; anomaly alerts; “compliance report generator” from SCP context Assumptions/dependencies: mapping SCP artifacts to regulatory frameworks; time-synced logs; retention and e-discovery policies
- Education and training: reproducible, safe “virtual-to-physical” lab exercises Sectors: universities, workforce development, MOOCs Tools/products/workflows: simulated SCP Servers for teaching; sandboxed protocols; “from literature to lab” assignments via protocol ingestion Assumptions/dependencies: access to simulator backends or shared teaching robots; pedagogical content; safe execution policies
- Lightweight research automation for SMEs and startups Sectors: small biotech/chem/materials labs, makerspaces Tools/products/workflows: SCP Server SDK for commodity instruments; template workflows (PCR, cell culture, titrations); managed Hub service Assumptions/dependencies: availability of low-cost device adapters; hosted Hub tiers; minimal IT overhead; vendor-neutral drivers
- Data-as-a-service for scientific corpora and model-assisted literature review Sectors: knowledge management, meta-research, competitive intelligence Tools/products/workflows: literature retrieval, structured extraction, citation graphs; “evidence-to-protocol” suggestions Assumptions/dependencies: content licensing/compliance; continual ingestion; deduplication and quality assurance
Long-Term Applications
- Global web of autonomous scientific agents coordinating across dry–wet labs Sectors: cross-disciplinary science at scale (all sectors) Tools/products/workflows: interlinked Hubs; standardized experiment-flow exchanges; multi-agent planning across institutions Assumptions/dependencies: broad protocol adoption; interoperability with adjacent standards; governance bodies and trust frameworks
- Routine autonomous hypothesis generation and closed-loop discovery at scale Sectors: pharma, materials for energy/storage, catalysis, agri-biotech Tools/products/workflows: objective-driven campaigns (design–make–test–learn) with dynamic resource allocation; outcome-aware planning Assumptions/dependencies: robust scientific reasoning agents; validated surrogate models; robust safety and escalation policies
- Journals and funders mandating machine-executable protocols with provenance Sectors: academic publishing, funding agencies, meta-science Tools/products/workflows: “submit JSON protocol + data lineage” alongside manuscripts; automated reproducibility checks Assumptions/dependencies: publisher policy shifts; community tooling for authoring/validation; long-term archival infrastructure
- Marketplaces for instrument time and method services (“RaaS”: Research-as-a-Service) Sectors: instrument vendors, CRO networks, cloud labs Tools/products/workflows: SCP-listed capabilities with pricing and SLAs; automated scheduling; escrowed provenance and results delivery Assumptions/dependencies: liability frameworks; standardized SLAs; billing/quotas; insurance and safety certifications
- Digital twins of laboratories and network-level optimization Sectors: operations for large lab networks, national facilities Tools/products/workflows: virtual labs mirroring devices; simulation of schedules, failure modes, and protocol variants before execution Assumptions/dependencies: high-fidelity device models; telemetry standards; continuous synchronization with physical assets
- Safety and governance orchestration for agentic science Sectors: policy/regulation, biosecurity, responsible AI Tools/products/workflows: policy-aware planners; risk scoring; automated guardrails and audits; red-team sandboxes Assumptions/dependencies: codified policy libraries; regulatory alignment; auditable AI decision trails; third-party certification
- Rapid-response scientific networks (e.g., outbreak countermeasures, disaster materials) Sectors: public health, emergency management, defense R&D Tools/products/workflows: federated Hubs pooling assays, models, and instruments; prioritized scheduling; publishable, executable protocols Assumptions/dependencies: emergency data-sharing compacts; cross-border compliance; pre-validated assays and device kits
- Large-scale automated replication and meta-analysis of published results Sectors: academia, policy evaluation for research funding Tools/products/workflows: bulk ingestion of methods; standardized re-execution on shared infrastructure; bias/error detection analytics Assumptions/dependencies: access to datasets/materials; legal permissions; scalable compute/lab capacity; community incentives
- Continuous, self-optimizing R&D loops tied to pilot manufacturing Sectors: advanced materials, chemicals, bioprocessing Tools/products/workflows: SCP-bridged lab→pilot line feedback; process-parameter optimization; multi-objective trade-off exploration Assumptions/dependencies: plant integration and safety; OT/IT convergence; robust sensors/QC; change-control compliance
- Citizen science and home-lab kits with constrained, safe automation Sectors: education, DIY science, community bio/maker spaces Tools/products/workflows: “safe-mode” SCP Servers for micro-instruments; curated protocols; risk-aware guidance Assumptions/dependencies: affordable, safe hardware; strong safety policies; curated protocol libraries; local compliance
These applications leverage SCP’s core contributions—unified resource integration, orchestrated experiment lifecycle management, secure/federated architecture, and a large tool ecosystem—to reduce integration overhead, enable reproducibility, and connect heterogeneous dry–wet capabilities across institutions. Feasibility in each context depends on standardized schemas and drivers, IAM and governance alignment, validated models and instruments, and sustained community and institutional adoption.
Glossary
- A-Lab: An autonomous materials synthesis laboratory that integrates AI and robotics for experimental synthesis. "A-Lab~\cite{szymanski2023autonomous}, an autonomous materials synthesis laboratory, has demonstrated the power of AI-driven experimental autonomy"
- ADMET: A set of pharmacokinetic and toxicity properties (Absorption, Distribution, Metabolism, Excretion, Toxicity) used to evaluate drug candidates. "combines cheminformatics, ADMET prediction, structural biology, and molecular docking"
- asynchronous anomaly notifications: System alerts delivered without blocking execution to report detected irregularities during workflows. "This subsystem provides asynchronous anomaly notifications"
- audit trail: A persistent, chronological record of actions and decisions for traceability and accountability. "while maintaining a persistent audit trail"
- AutoDock Vina: A widely used molecular docking engine for predicting ligand-protein binding poses and affinities. "an AutoDock Vina-style docking engine"
- binding affinities: Quantitative measures of how strongly a ligand binds to a target, often in kcal/mol. "records the predicted binding affinities"
- binding pockets: Cavities on a protein surface where small molecules can bind during docking or interactions. "identify putative binding pockets with fpocket"
- cheminformatics: The application of computational methods and data analysis to chemical problems and molecular data. "combines cheminformatics, ADMET prediction, structural biology, and molecular docking"
- closed-loop control: A feedback-driven strategy where outputs are monitored to iteratively adjust subsequent actions. "closed-loop control of robotic laboratory instruments"
- conflict detection: Identification of incompatible resource demands or constraints within planned workflows. "performs conflict detection and resource forecasting"
- cross-coupling reactions: Chemical reactions that form carbon–carbon bonds by coupling two molecular fragments, often catalyzed by metals. "optimizing cross-coupling reactions"
- data-governance policies: Institutional rules and processes that regulate data access, usage, security, and compliance. "integration with institutional data-governance policies"
- dependency structure: The graph of prerequisite relationships among tasks within a plan or workflow. "together with dependency structure, expected latency, experimental risk, and cost estimates"
- ELN: An Electronic Lab Notebook used to record, manage, and export laboratory protocols and data. "a lab SOP exported from an ELN"
- experiment context: A structured metadata object that captures identifiers, goals, types, storage URIs, and configurations for an experiment. "SCP defines a first-class experiment context that records a persistent experiment identifier, experiment type (dry, wet, or hybrid), high-level goals, data storage URIs, and configuration parameters"
- experiment-flow API: An interface for generating, recommending, and invoking multi-step experimental workflows programmatically. "The intelligent orchestration mechanism is exposed via the experiment-flow API"
- failover: Automatic switching to backup resources or strategies when failures occur to maintain continuity. "These reports inform global scheduling and failover decisions made at the Hub level"
- fpocket: A computational tool used to detect and score potential ligand-binding pockets in protein structures. "identify putative binding pockets with fpocket"
- high-performance computing cluster: A distributed computing system that provides large-scale computational resources for simulations or analysis. "simulation on a remote high-performance computing cluster"
- hub--and--spoke architecture: A design pattern where a central hub coordinates multiple peripheral nodes (spokes). "SCP adopts a hub--and--spoke architecture"
- intent-analysis models: AI models that infer user goals from natural-language instructions to generate candidate task graphs. "uses AI-driven intent-analysis models to translate natural-language instructions into a set of candidate task graphs"
- intent recognition module: A system component that parses inputs to identify tasks and constraints for planning. "The SCP Hub’s intent recognition module parses the input to identify high-level tasks and constraints"
- LD\textsubscript{50}: The median lethal dose, a toxicity metric indicating the dose required to kill 50% of a test population. "the pred_molecule_admet tool predicts the LD\textsubscript{50} toxicity metric"
- ligands: Small molecules that bind to a target protein in docking and biochemical interactions. "both ligands and receptor must be converted into PDBQT format"
- Model Context Protocol (MCP): A standard for connecting AI models to tools and data sources via uniform interfaces. "Existing standards (e.g. the Model Context Protocol) do not fully resolve these issues"
- OAuth2.1: A modern authorization framework version enabling secure, fine-grained access control for users and experiments. "OAuth2.1-based authentication and authorization for experiments and users"
- organocatalysts: Organic molecules that catalyze chemical reactions without requiring metal centers. "from organocatalysts to optimizing cross-coupling reactions"
- PDB: The Protein Data Bank, a repository of 3D structural data of proteins and nucleic acids. "Using PDB ID~6vkv as the target"
- PDBFixer: A tool that repairs missing or inconsistent regions in PDB structures prior to modeling or docking. "(iii) repair missing or inconsistent regions using PDBFixer"
- PDBQT: A file format used by AutoDock tools that includes atom types and partial charges for docking. "converted into PDBQT format"
- provenance: The record of data origins, transformations, and lineage for reproducibility and auditing. "including experiment registration, planning, execution, and provenance tracking"
- QED (Quantitative Estimate of Drug-likeness): A scalar measure indicating how drug-like a molecule is based on multiple medicinal chemistry attributes. "QED (Quantitative Estimate of Drug-likeness) score"
- receptor protein: The target macromolecule prepared for docking to evaluate ligand binding. "prepares the receptor protein for docking"
- SMILES: A line notation that encodes molecular structures as text strings. "encoded in SMILES format"
- SOP: A Standard Operating Procedure that formalizes step-by-step methods in laboratory practice. "a lab SOP exported from an ELN"
- spectroscopy: Analytical techniques that measure interactions between matter and electromagnetic radiation. "a spectroscopy tool"
- Thoth Server: An SCP-compatible protocol understanding node that converts free-form or high-level requests into structured experimental protocols. "the Thoth Server is invoked as a specialized SCP service node to generate the detailed laboratory protocol"
- Thoth-OP Server: An SCP execution-planning node that decomposes protocols into device-specific command sequences. "the Thoth-OP Server decomposes the structured protocol into atomic operations"
- wet-lab device integration: The inclusion of physical laboratory equipment as addressable tools within the same protocol as computational resources. "Wet-lab device integration. SCP extends the notion of a tool to cover real laboratory equipment"
- workflow orchestration: Coordinated planning, scheduling, and supervision of multi-step workflows across tools and agents. "Intelligent workflow orchestration."
- workflow specification: A formal, machine-readable description of steps, parameters, outputs, and dependencies in an experiment. "fine-grained, protocol-level workflow specification"
Collections
Sign up for free to add this paper to one or more collections.