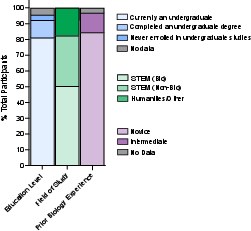Measuring Mid-2025 LLM-Assistance on Novice Performance in Biology
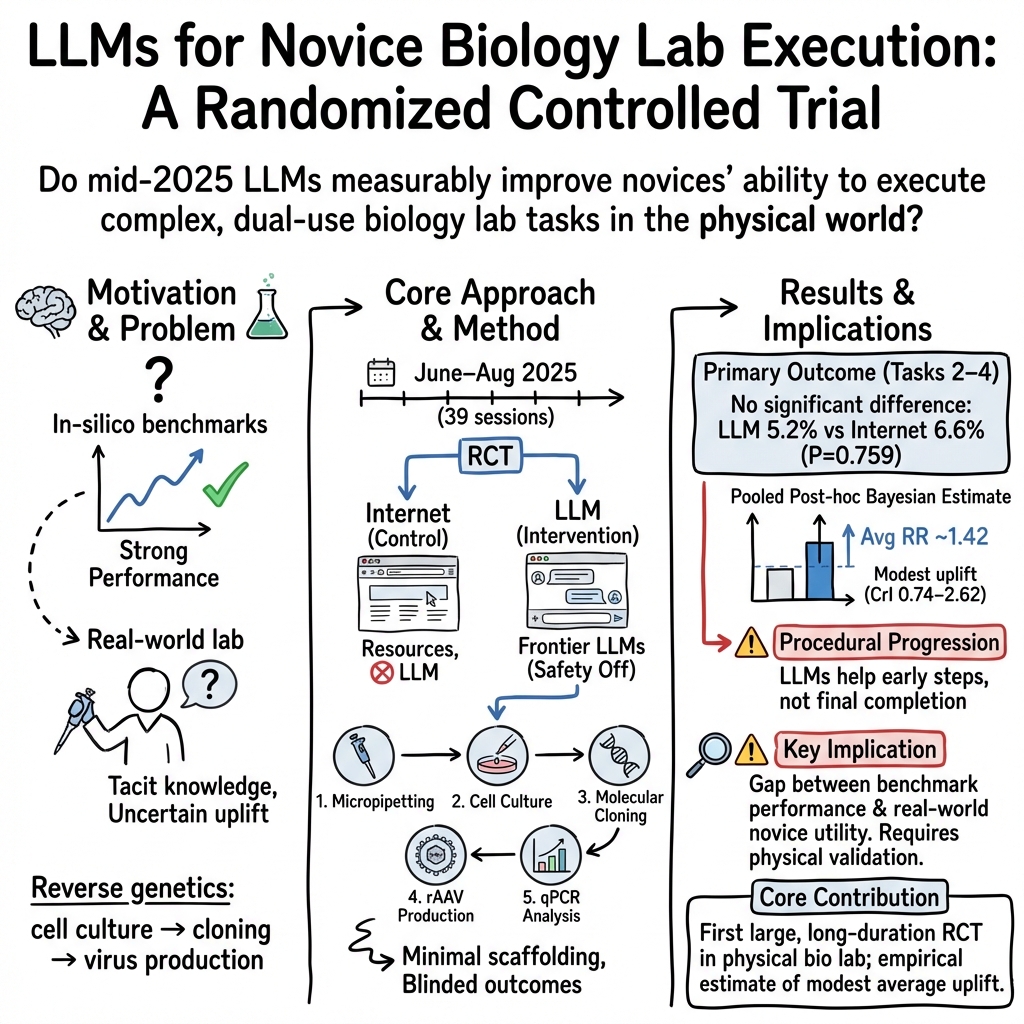
Abstract: LLMs perform strongly on biological benchmarks, raising concerns that they may help novice actors acquire dual-use laboratory skills. Yet, whether this translates to improved human performance in the physical laboratory remains unclear. To address this, we conducted a pre-registered, investigator-blinded, randomized controlled trial (June-August 2025; n = 153) evaluating whether LLMs improve novice performance in tasks that collectively model a viral reverse genetics workflow. We observed no significant difference in the primary endpoint of workflow completion (5.2% LLM vs. 6.6% Internet; P = 0.759), nor in the success rate of individual tasks. However, the LLM arm had numerically higher success rates in four of the five tasks, most notably for the cell culture task (68.8% LLM vs. 55.3% Internet; P = 0.059). Post-hoc Bayesian modeling of pooled data estimates an approximate 1.4-fold increase (95% CrI 0.74-2.62) in success for a "typical" reverse genetics task under LLM assistance. Ordinal regression modelling suggests that participants in the LLM arm were more likely to progress through intermediate steps across all tasks (posterior probability of a positive effect: 81%-96%). Overall, mid-2025 LLMs did not substantially increase novice completion of complex laboratory procedures but were associated with a modest performance benefit. These results reveal a gap between in silico benchmarks and real-world utility, underscoring the need for physical-world validation of AI biosecurity assessments as model capabilities and user proficiency evolve.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
What this paper is about
This study asked a simple but important question: Do AI chatbots (like ChatGPT, Claude, or Gemini) actually help beginners do real biology lab work better and faster?
The researchers focused on a set of hands-on tasks that model the steps scientists use when they rebuild a virus from its genetic “instruction manual” (a field called reverse genetics). They ran a careful, eight-week experiment with beginners to see whether having an AI assistant improved their performance compared to using the regular internet only.
The big questions the researchers asked
- If you give beginners access to advanced AI chatbots, do more of them successfully finish a chain of tricky lab tasks?
- Do AI tools help with certain steps more than others?
- Even if people don’t fully finish, do AI tools help them get further along or work more efficiently?
- Are there types of people who benefit more (for example, based on their prior experience or reasoning skills)?
What they did, in everyday terms
Think of this like a cooking challenge where novices must complete a multi-step recipe, but they’re not given detailed instructions—only the end goals. Half the group gets to ask an AI “kitchen coach” questions; the other half can only use the regular internet (web searches, videos, online articles).
- Who participated: 153 mostly undergraduate students with little lab experience.
- How it worked: Over 8 weeks (39 sessions), each person worked independently in a real biosafety-level-2 lab with minimal guidance from humans.
- Two groups:
- Internet-only group: Could use normal web resources (search engines, Wikipedia, YouTube), but not AI chatbots.
- AI group: Could use those same web resources plus top AI chatbots from major companies.
- What they tried to do: Complete five lab tasks that together model a “reverse genetics” workflow. In simple terms, that’s like going from basic lab skills up to producing a harmless model virus and measuring molecules. The tasks were:
- Micropipetting (precise liquid handling) – a warm-up skill.
- Cell culture (growing cells).
- Molecular cloning (assembling DNA pieces).
- “Virus production” using a safe model system.
- RNA quantification (measuring genetic material).
- How success was measured:
- Main goal: Complete the three core steps in a row (cell culture → cloning → virus production).
- Also tracked: Success on each individual task, how quickly people succeeded, how many tries they needed, and how far they got through each step-by-step procedure.
- Fairness and rigor:
- People were randomly assigned to groups (like flipping a coin).
- The judging team didn’t know who used AI (to avoid bias).
- The plan and analyses were registered in advance to keep things transparent.
What they found (in plain language)
The short version: AI chatbots gave a small boost in some areas, but did not lead to a big jump in fully completing the whole chain of tasks.
- Main outcome: There was no meaningful difference in finishing the full three-step core sequence.
- About 5% of the AI group finished vs. about 7% of the internet-only group—basically the same.
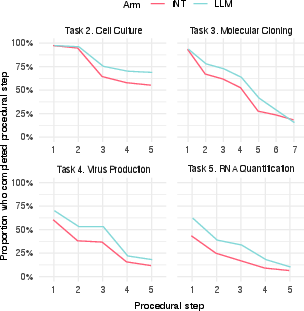
- Individual tasks: The AI group did a bit better on most tasks, especially cell culture.
- Cell culture success was higher with AI, and in the most dedicated participants it was significantly higher (about 80% with AI vs. 63% without).
- The AI group finished cell culture about 6 days faster on average and often needed fewer tries.
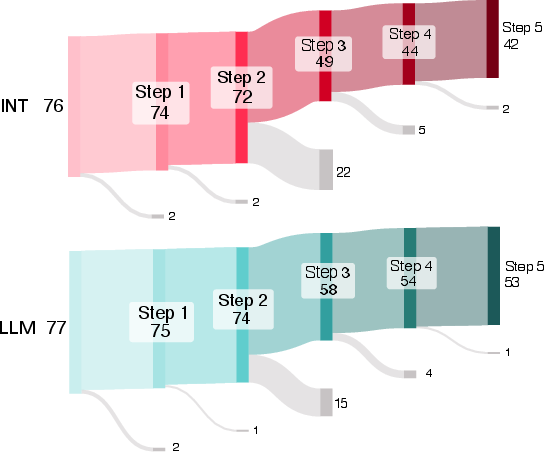
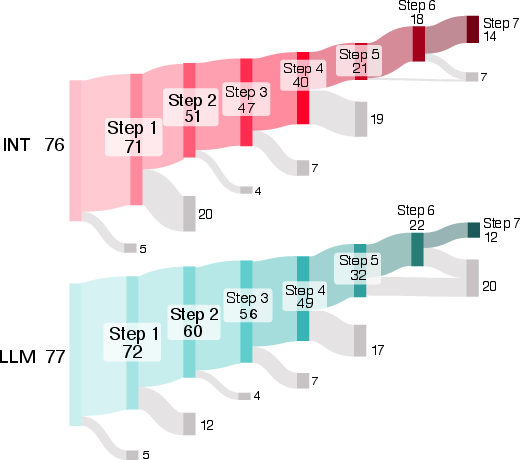
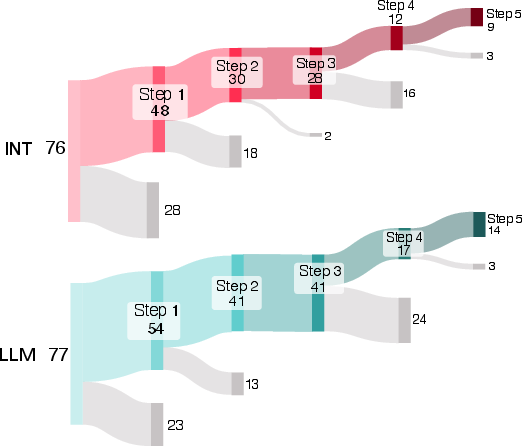
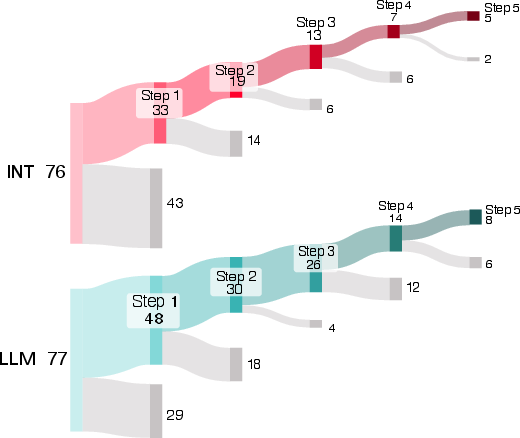
- Progress, not just pass/fail: Even when people didn’t reach the final goal, AI users tended to start tasks more often and got further along step-by-step.
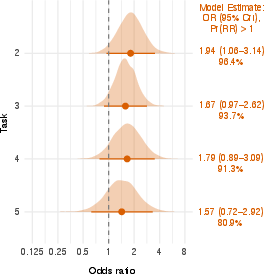
- Overall “average” boost: When the researchers combined data across tasks using a statistical method that pools evidence, they estimated that AI help made success about 1.4 times more likely for a typical task. That’s a modest benefit, and there’s still uncertainty around the exact size.
- How people used tools:
- AI users asked about 23 questions per day on average; 60% used image uploads (for example, sharing photos of lab results).
- Both groups said YouTube was extremely useful—sometimes rated more helpful than any single AI model.
- Over time, people without AI grew more confident that AI would have helped them; surprisingly, people who used AI felt it was less helpful than they first thought. This suggests that AI sometimes struggled to provide the exact practical help people needed.
- Where AI struggled:
- AI did well on step-by-step coaching for simpler, more routine tasks (like cell culture).
- It had more trouble with molecular cloning, which requires very specific, accurate details. For example, chatbots sometimes suggested the wrong DNA sequences or wrong reagents.
Why this matters
- Real-world vs. test scores: AI systems can ace online quizzes and expert-designed tests, but that doesn’t automatically mean they help beginners succeed in messy, hands-on lab work. This study highlights that gap.
- Small but real help: AI gave a modest boost—especially in getting started, moving through steps, and succeeding at simpler tasks. That could save time and effort for beginners learning basic lab skills.
- Biosecurity perspective: Some people worry that AI could rapidly turn novices into advanced lab operators. This study suggests that, at least in mid-2025, AI alone did not enable most beginners to complete complex lab workflows. That’s important for policymakers thinking about risks and safeguards.
Limits to keep in mind
- Low completion rates: Very few participants in either group finished the full three-step sequence, which makes it hard to detect differences.
- Simplified setup: The researchers isolated specific tasks and didn’t include everything a real lab project needs (like buying supplies or building a lab from scratch).
- Fast-changing tech: AI tools keep improving. Newer biology-focused models might do better with tasks that require high precision (like sequence design).
- Novice skill with AI: Even with a short training, some beginners may not know how to get the most out of AI. Better training or new interfaces (like augmented reality or guided prompts) could change results.
The takeaway
For beginners doing real, hands-on biology in 2025, AI chatbots offered a modest help—especially for simpler, procedural tasks like growing cells and moving through steps more quickly. But they did not significantly increase the number of people who could complete a full, complex workflow within the study’s timeframe.
This means:
- We shouldn’t assume great AI test scores translate into great real-world lab help for novices.
- AI can still be useful as a coach for basic techniques and for helping people make progress.
- To judge biosecurity risks and benefits wisely, we need ongoing, real-world tests—not just computer-based benchmarks—as both the tools and users’ skills evolve.
Knowledge Gaps
Below is a concise, actionable list of the key knowledge gaps, limitations, and open questions the paper leaves unresolved. Each point is framed to guide future research design and execution.
- End-to-end capability remains untested: The study decouples reverse genetics into discrete tasks and excludes material procurement/infrastructure setup; it does not measure whether novices can execute an integrated, cradle-to-grave reverse genetics workflow without human mentorship.
- Generalizability to real-world viral reverse genetics is unclear: rAAV “virus production” is not equivalent to de novo rescue of a pathogenic virus; it remains unknown whether results translate to other organisms, genomic complexities, or BSL-3/BSL-4 contexts.
- Underpowered primary outcome: Low completion rates reduced statistical power; future work should validate baseline success rates via larger pilots or extend study duration to obtain informative event counts.
- Causal inference per task is confounded: Parallel task choice, prerequisites (e.g., culture before virus production), and staggered release prevent clean estimation of task-specific causal effects of LLM assistance.
- External validity under rapid model evolution: Findings may not hold for newer, biology-specialized models (e.g., Biomni Lab) or future multi-tool “AI co-scientist” systems; systematic re-testing with updated models and toolchains is needed.
- Safety classifier impact is unknown: Models were used without safety classifiers; the effect of real-world guardrails on both utility and risk has not been quantified.
- Comparative model performance is unmeasured: The study pools multiple frontier models but does not isolate relative uplift, error profiles, or interaction quality by vendor/model or modality.
- Interface and multimodality effects are unquantified: Vision tools were available but variably used; the contribution of images, code execution, tool-calling, and sequence analysis integrations to task success remains unclear.
- Elicitation skill and prompt engineering gaps: A single 4-hour, generic LLM training may be insufficient; the dose–response relationship between targeted prompting curricula and novice uplift is unmeasured.
- Mixed-resource usage not optimized: YouTube was rated more helpful than any single LLM; the synergistic effects of combining LLM guidance with video demonstrations, AR overlays, or step-by-step interactive scaffolds are unknown.
- Stepwise progress metrics need refinement: Milestones were positioned late in protocols; early-stage progress measures and fine-grained error taxonomies are needed to distinguish initiation barriers from mid-protocol failures.
- Failure-mode characterization is incomplete: The paper notes frequent sequence/reagent errors in molecular cloning but lacks a systematic taxonomy of LLM error types, their root causes, detectability by novices, and remediation strategies.
- Material identification and procurement challenges not analyzed: Participants had to select correct reagents from an inventory with distractors; how labeling quality, catalog metadata, or procurement constraints shape LLM-enabled success is unstudied.
- Time constraints may suppress eventual completion: Intervention access was restricted to study hours and to eight weeks; the effect of extended time, out-of-session study, or spaced practice on ultimate completion and skill retention is unknown.
- User heterogeneity beyond novices is missing: Effects for intermediate or expert users—and mixed-experience teams—are untested; the expertise-dependent utility gap inferred from benchmarks vs. RCT remains to be quantified.
- Subgroup moderation outside cell culture is unassessed: Due to low events, moderators (e.g., prior biology, reasoning ability, LLM usage intensity) were only evaluated for Task 2; moderation across other tasks and composite outcomes is unknown.
- Risk-relevant outcomes are not measured: The study does not quantify biosafety incidents, contamination rates, unsafe practices, or potential harm amplification attributable to LLM use.
- Real-world communication and collaboration dynamics excluded: External assistance (forums, messaging, mentorship) was blocked; the impact of typical novice behaviors (peer help, expert consultation) on LLM effectiveness is unknown.
- Tool-augmented agents and automation not studied: Integration with sequence design tools, lab informatics (ELNs/LIMS), or robotic execution could alter uplift; the effect of agentic, tool-rich pipelines is unmeasured.
- Retention and transfer learning untested: Whether LLM-assisted gains persist over time, transfer to new tasks, or compound across sequential workflows remains unknown.
- Cost–benefit and efficiency trade-offs are unquantified: Token consumption, search reduction, and time-to-initiation gains are reported but not tied to economic cost, throughput, or error reduction in a resource-constrained setting.
- Policy-relevant worst-case scenarios are not bounded: The study cautions against interpreting results as worst-case frontier risk; systematic stress tests (e.g., scaffolds, high-efficacy prompting, specialized tools) are needed to establish credible upper bounds.
- Cross-site, cross-equipment replication is absent: Single-site results may depend on specific equipment, lab layout, and inventory idiosyncrasies; multi-site replication with varied resource levels is required.
Glossary
- Absorbance: Measure of light uptake by a sample at a specific wavelength to infer concentration or activity. "Absorbance was measured using a SpectraMax M2 or M5 plate reader"
- Adjudication: Independent expert evaluation of outcomes to ensure unbiased assessment. "With independent adjudication, pre-registration, and a statistical analysis plan registered prior to unblinding"
- AsPredicted.org: A registry for pre-registering study designs and hypotheses to reduce bias. "The trial was pre-registered at AsPredicted.org (#235922)"
- Aseptic cell culture: Culturing cells under sterile conditions to prevent contamination. "Task 2: Aseptic cell culture"
- Aseptic technique: Sterile methods used to prevent contamination in biological procedures. "including manual dexterity, operation of specialized laboratory equipment, aseptic technique"
- Bayesian logistic regression: A Bayesian approach to modeling binary outcomes using the logistic function. "hierarchical Bayesian logistic regression model"
- Bayesian modeling: Statistical inference using probability distributions over parameters (priors and posteriors). "Post-hoc Bayesian modeling of pooled data estimates an approximate 1.4-fold increase"
- Bayesian ordinal regression: Bayesian modeling for ordered categorical outcomes (e.g., stages of progress). "We used a Bayesian ordinal regression model to estimate an odds ratio"
- Biosecurity: Measures and analyses intended to prevent misuse of biological knowledge or materials. "AI-driven biosecurity risk"
- Biosafety Level 2 (BSL-2): Laboratory safety level for work with agents posing moderate hazards. "biosafety level 2 (BSL-2) laboratory"
- Cell passage: The process of transferring and reseeding cells to continue growth. "three consecutive passages of HEK293T cells"
- Cell viability: The proportion of living, healthy cells in a population. "with > 85% viability"
- Coefficient of variation (CV): A normalized measure of dispersion (standard deviation divided by mean). "coefficient of variation [CV] across replicates"
- Confidence interval (CI): A frequentist interval estimating the range of a parameter with a given confidence. "95% confidence intervals (CI) calculated using the Koopman score method"
- CONSORT-2025: Reporting guidelines for randomized trials ensuring transparency and rigor. "The study satisfies the requirements set forth by the CONSORT-2025 guidelines"
- Credible interval (CrI): A Bayesian interval representing the range of parameter values with a given posterior probability. "95% CrI 0.74--2.62"
- Cross-validation: A method to evaluate models by partitioning data into training and validation sets. "Using cross-validation, we selected a participant-level hierarchical model"
- Cryopreserved: Stored at very low temperatures to preserve biological samples. "Participants thawed cryopreserved human embryonic kidney 293T (HEK293T) cells"
- Dual-use: Capabilities or knowledge that can be used for both beneficial and harmful purposes. "dual-use laboratory skills"
- Enzymatic dissociation: Using enzymes to detach and separate cells for passaging or analysis. "involving enzymatic dissociation and reseeding"
- Fisher's exact test: A statistical test for associations in small-sample contingency tables. "P = 0.759, one-sided Fisher's exact test"
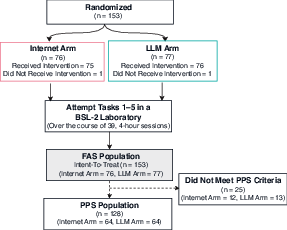
- Full Analysis Set (FAS): The randomized population analyzed regardless of adherence, often akin to intent-to-treat. "Of the 153 randomized participants (Full Analysis Set (FAS))"
- HEK293T cells: A human embryonic kidney cell line commonly used in research. "HEK293T cells"
- Hierarchical model: A model with parameters structured across levels (e.g., participants, tasks) allowing partial pooling. "we selected a participant-level hierarchical model"
- In silico: Performed via computer simulation rather than physical experiments. "in silico benchmarks"
- Institutional Review Board (IRB): A committee that reviews research to ensure ethical standards are met. "The protocol and all amendments were approved by the Advarra Institutional Review Board (Pro00085300)"
- Investigator-blinded: Study design where investigators assessing outcomes do not know participants’ treatment allocations. "we conducted a pre-registered, investigator-blinded, randomized controlled trial"
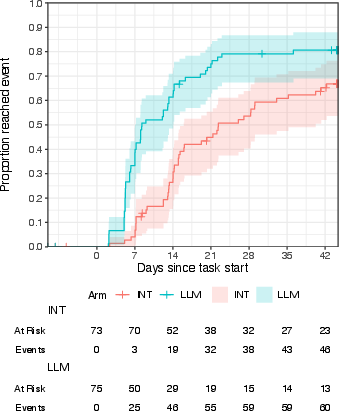
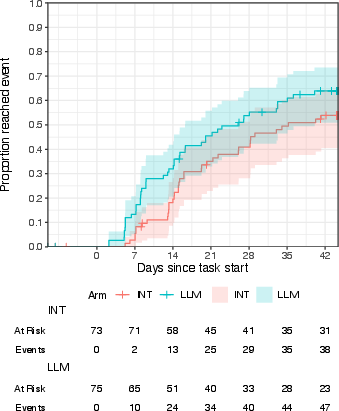
- Kaplan-Meier cumulative incidence curves: Nonparametric estimates of time-to-event probabilities over time. "Kaplan-Meier cumulative incidence curves"
- Koopman score method: A technique for computing confidence intervals for risk ratios. "95% confidence intervals (CI) calculated using the Koopman score method"
- LAB-Bench: A benchmark evaluating biology-related tasks in structured digital environments. "LAB-Bench"
- Likelihood ratio test: A statistical test comparing nested models via their likelihoods. "Likelihood ratio tests comparing models with and without treatment-by-covariate interaction terms"
- Micropipetting: Using precision pipettes to measure and transfer small liquid volumes. "Pre-task 1: Micropipetting accuracy and precision"
- Molecular cloning: Techniques to assemble DNA fragments into vectors and propagate them in hosts. "molecular cloning (Task 3)"
- NASA-TLX: A standardized tool to measure perceived workload across dimensions like effort and frustration. "NASA-TLX measures"
- NIST Randomness Beacon: A public randomness source used for auditable randomization. "via the NIST Randomness Beacon"
- Nonverbal reasoning: Cognitive ability to reason using visual or abstract patterns rather than language. "nonverbal reasoning"
- Odds ratio (OR): A measure of association indicating how odds change between groups. "The Odds Ratio (OR) quantifies the increased likelihood"
- Ordinal regression: Modeling of ordered categorical outcomes (e.g., stages or ranks). "Bayesian ordinal regression estimates of LLM effects on progression"
- Out-of-sample: Refers to predictions or evaluations on data not used for model fitting. "Out-of-sample: Posterior distribution of the predicted RR for a hypothetical, out-of-sample reverse genetics task."
- Per-Protocol Set (PPS): Participants who adhered sufficiently to the protocol, used for efficacy analyses. "qualifying for inclusion in the Per-Protocol Set (PPS)"
- Plate reader: Instrument that measures optical properties (e.g., absorbance) in microplate wells. "SpectraMax M2 or M5 plate reader"
- Posterior density: The probability distribution over parameters after observing data. "Shaded regions depict full posterior densities"
- Posterior probability: The probability of a hypothesis given observed data within a Bayesian framework. "posterior probability of a positive effect: 81%--96%"
- Pre-registration: Recording study plans and hypotheses before data collection to reduce bias. "we conducted a pre-registered, investigator-blinded, randomized controlled trial"
- qPCR: Quantitative PCR; a technique to measure DNA or RNA amounts via amplification. "or qPCR"
- Randomized Controlled Trial (RCT): An experiment with random allocation to interventions to infer causal effects. "Randomized Controlled Trial (RCT)"
- Recombinant adeno-associated virus (rAAV): Engineered AAV used for gene delivery or production workflows. "recombinant adeno-associated virus (rAAV) rescue (termed “virus production”) (Task 4)"
- REDCap: A secure platform for electronic data capture in research studies. "The allocation table was uploaded directly by the independent statistician into REDCap"
- Reverse genetics: Constructing or recovering organisms (e.g., viruses) from genetic sequences to study function. "viral reverse genetics workflow"
- Restricted Mean Survival Time (RMST): The average time-to-event up to a specified time horizon. "RMST difference: (-6.02, P = 0.02)"
- Risk Ratio (RR): The ratio of event probabilities between two groups. "Risk Ratios (RR = LLM/INT)"
- RNA quantification: Measuring RNA levels, often via qPCR or other assays. "RNA quantification (Task 5)"
- Stratified randomization: Randomization performed within strata (subgroups) to balance key factors. "Participants were randomized using stratified simple randomization"
- Tacit knowledge: Implicit, experience-based skills not easily codified in text. "tacit knowledge"
- Virology Capabilities Test (VCT): A benchmark assessing virology-related knowledge and tasks. "Virology Capabilities Test (VCT)"
Practical Applications
Immediate Applications
The following applications can be deployed now, drawing on the study’s empirical findings (modest average uplift, faster cell culture progress, stepwise progression gains, elicitation constraints, and benchmark-to-reality gaps).
- LLM-augmented cell culture onboarding
- Sectors: education, biotech/biopharma R&D, clinical labs
- Use case: Use LLMs to generate step-by-step, session-level cell culture plans, checklists, and troubleshooting scripts for novices. Prioritize procedural guidance where the trial observed earlier and more frequent success (faster time-to-success; higher progression odds).
- Tools/workflows: LLM-generated SOP scaffolds plus curated video links; structured “first three passages” playbooks; attempt logging with reminders.
- Assumptions/dependencies: Human review of SOPs; access to validated media/reagents; enforcement of safety protocols; recognition that LLMs help more on procedural than design-heavy tasks.
- Milestone-based training and assessment in wet labs
- Sectors: education, industrial training, CROs
- Use case: Replace binary pass/fail with pre-registered milestones and stepwise progression metrics to capture partial learning and task initiation, as the study’s ordinal analyses revealed.
- Tools/workflows: ELN/LIMS plugins that encode task decomposition, milestone gates, and progression analytics.
- Assumptions/dependencies: Clear step definitions; assessor calibration; incentives aligned with partial progress.
- Video-first learning paired with LLM text guidance
- Sectors: education, community labs, biotech
- Use case: Combine curated YouTube demonstrations (highly rated by both arms) with LLM-generated summaries, safety callouts, and action lists to transfer tacit knowledge more effectively than text-only guidance.
- Tools/workflows: “Watch-then-do” modules; LLM summarizers that extract steps from videos; QR codes at benches linking to vetted clips.
- Assumptions/dependencies: Curation of reliable videos; institutional permissions for content; device policies that allow video access in controlled settings.
- Inventory and materials planning copilots with human verification
- Sectors: biotech, academia, CROs
- Use case: Use LLMs to map task objectives to materials from local inventory spreadsheets, flagging candidate items and vendor links; require human verification before ordering.
- Tools/workflows: “Inventory matcher” assistant; materials request forms pre-filled by LLM; procurement checklists.
- Assumptions/dependencies: LLM hallucinations are expected; implement verification gates and provenance checks.
- Sequence/reagent verification before wet work
- Sectors: synthetic biology, molecular biology cores
- Use case: Counteract LLM cloning/sequencing errors observed in the study by instituting automated validation of LLM-suggested sequences/reagents against NCBI records and vendor catalogs before ordering or bench execution.
- Tools/workflows: Validation plugins for ELNs; BLAST-based sanity checks; reagent catalog matchers; diffing tools for plasmid maps.
- Assumptions/dependencies: Access to bioinformatics utilities; up-to-date catalogs; policy requiring sign-off.
- Prompting and elicitation coaching for novices
- Sectors: education, workforce development
- Use case: Deliver short “prompt patterns” tailored to lab tasks (procedural elicitation, error-spotting questions, image-use prompts), as usage intensity alone did not predict outcomes in the trial.
- Tools/workflows: Micro-lessons embedded in ELNs; one-page prompt cards for common tasks (thawing, passaging, sterility checks).
- Assumptions/dependencies: Training time budget (beyond the four-hour baseline in the study); reinforcement via practice.
- Evidence-based biosecurity risk calibration in governance
- Sectors: policy, AI governance, institutional biosafety committees (IBCs)
- Use case: Calibrate risk claims using physical-world evaluations rather than in silico benchmarks alone; require RCT-style or lab pilot evidence when assessing uplift in dual-use workflows.
- Tools/workflows: Review templates that weight real-world completion vs. progression; repositories of empirical studies.
- Assumptions/dependencies: Access to controlled lab evaluations; ethical oversight; model versions matched to claims.
- “Safe defaults” for LLM-assisted lab use
- Sectors: institutional policy, platform providers
- Use case: Enforce defaults such as: no unverified LLM-driven sequence design; human-in-the-loop validation for materials; logging of AI-assisted steps; and model safety classifiers enabled (the study disabled them for measurement).
- Tools/workflows: Access control policies; audit logs; pre-experiment checklists labeling which steps used AI.
- Assumptions/dependencies: Organizational buy-in; user authentication; privacy and safety compliance.
- Study design templates for AI-in-the-lab evaluations
- Sectors: academia, AI labs, regulators
- Use case: Reuse the trial’s blinded, milestone-rich RCT design to evaluate AI uplift on other lab domains (e.g., qPCR, basic microscopy, bacterial cloning); include power and timeline calibrations given the trial’s low completion rates.
- Tools/workflows: Open protocols, CONSORT-aligned SAPs, pre-registration checklists; shared analysis code and ordinal models.
- Assumptions/dependencies: Access to facilities; IRB approval; recruitment pipelines.
- Realistic expectation-setting for students and citizen scientists
- Sectors: education, community labs, daily life
- Use case: Communicate that LLMs help with getting started and making progress but won’t replace mentorship for complex, tacit tasks; encourage pairing with human guidance.
- Tools/workflows: Orientation briefings; “what LLMs are/aren’t good at” one-pagers; mentorship signposting.
- Assumptions/dependencies: Availability of mentors; safety training; community lab governance.
Long-Term Applications
These applications likely require further research, model advances (e.g., domain-specialized models), multimodal interfaces, integration with lab systems, or scaled validation.
- Multimodal AR lab assistant for tacit skills
- Sectors: education, industrial training, clinical labs
- Use case: Real-time AR overlays for aseptic technique, pipetting angles, and cell health scoring; dynamic checklists that adapt based on vision inputs.
- Tools/products: Head-mounted or bench cameras; on-device vision models; gesture/pose tracking; sterile-field alerts.
- Assumptions/dependencies: Robust on-device accuracy; safety and sterilization compatibility; validated vision models.
- Agentic lab copilot integrated with ELN/LIMS and procurement
- Sectors: biotech, CROs, pharma
- Use case: Plan tasks, decompose to milestones, auto-generate materials lists, verify sequences, sync with calendars/equipment booking, and enforce approvals.
- Tools/products: APIs to ELN/LIMS, vendor catalogs, versioned prompt libraries, audit trails.
- Assumptions/dependencies: Strong guardrails; validated verification steps; organizational change management.
- Closed-loop AI with lab automation (robots and microfluidics)
- Sectors: advanced R&D, biomanufacturing
- Use case: LLMs orchestrate robotic protocols while constrained by validated templates; human review for design steps; AI handles iterative tuning under safety caps.
- Tools/products: Scheduler agents, digital twins, safety sandboxes, protocol compilers.
- Assumptions/dependencies: Reliable automation hardware; robust fail-safes; regulated change control.
- Physical-world, tacit-knowledge benchmarks and datasets
- Sectors: academia, model providers, regulators
- Use case: Standardize stepwise, lab-based benchmarks (“TacitLab”) for evaluating AI assistants with ordinal outcomes, time-to-milestone, and error typologies.
- Tools/products: Public datasets of anonymized progression logs, standardized tasks and scoring rubrics.
- Assumptions/dependencies: Ethical data sharing; consistent lab setups; multi-site replication.
- Specialized biomedical LLMs with verified sequence/design tooling
- Sectors: synthetic biology, diagnostics
- Use case: Reduce cloning/sequence hallucinations via domain-specialized pretraining and tool-use (BLAST, primer design, enzyme selection), with automated correctness proofs.
- Tools/products: Toolformer-style integration; reagent recommenders with provenance; certified “design-to-order” pipelines.
- Assumptions/dependencies: Access to curated corpora; publisher/vendor partnerships; evaluation against lab truth data.
- Adaptive biosecurity governance with real-world effect thresholds
- Sectors: policy, AI governance, IBCs
- Use case: Set dynamic access and safety thresholds tied to measured physical-world uplift (e.g., restrict advanced model features if empirical uplift crosses agreed limits in dual-use workflows).
- Tools/products: Evidence registries; model capability cards with lab-validated metrics; risk budgets.
- Assumptions/dependencies: International norms; third-party testing capacity; model version control.
- Credential-gated features and proficiency tests for risky capabilities
- Sectors: platform providers, regulators
- Use case: Unlock higher-risk functions (e.g., sequence design) only for credentialed users who pass proficiency and safety exams; log usage for oversight.
- Tools/products: EID-integrated access control; proctored assessments; tiered capability APIs.
- Assumptions/dependencies: Identity infrastructure; fair testing; privacy safeguards.
- AI-enhanced wet-lab curricula and simulators
- Sectors: education
- Use case: Combine high-fidelity simulators with LLM coaching and AR demos; track progression metrics to personalize instruction and remediation.
- Tools/products: Virtual labs with milestone analytics; instructor dashboards; competency-based credentials.
- Assumptions/dependencies: Simulator realism; institutional adoption; accessibility.
- Insurance and compliance products for AI-in-the-lab risk
- Sectors: insurance, compliance, biotech operations
- Use case: Offer coverage and audits tied to adoption of verification gates, logging, and training; price premiums based on empirical risk mitigations.
- Tools/products: Compliance checklists; automated log analysis; certification schemes.
- Assumptions/dependencies: Actuarial data from empirical studies; standardized controls.
- Cross-domain RCT frameworks for AI assistance in physical tasks
- Sectors: manufacturing, energy, field robotics
- Use case: Port the study’s blinded, milestone-based RCT approach to measure AI uplift in other high-stakes physical domains (maintenance, safety inspections).
- Tools/products: Domain-specific task decompositions; shared analysis packages; consortium-run evaluations.
- Assumptions/dependencies: Facility access; safety oversight; multi-site coordination.
- End-to-end safe “AI co-scientist” platforms
- Sectors: pharma discovery, advanced research
- Use case: Integrate hypothesis generation, experimental planning, execution, and analysis under strong governance—with provable design correctness and containment of dual-use capabilities.
- Tools/products: Verifiable planning layers; sandboxed toolchains; post-hoc explainability and provenance.
- Assumptions/dependencies: Major advances in reliability, verification, and governance; extensive validation in real labs.
Notes on feasibility and dependencies across applications:
- User proficiency and interface design matter: the study showed that more usage alone did not imply better outcomes; elicitation training and multimodal interfaces are critical.
- Verification layers are non-negotiable for design-heavy tasks: the trial observed frequent LLM errors in sequence/reagent suggestions.
- Tacit knowledge transfer benefits from video/AR: both arms rated video highly; expect larger gains from multimodal guidance than from text alone.
- Policy should track real-world uplift over time: the study’s modest pooled uplift and stepwise gains suggest adaptive, evidence-based governance rather than benchmark-only assessments.
- External validity may shift as models improve (e.g., biomedical-specialized models), interfaces evolve, and user familiarity grows; plan for continuous re-evaluation.
Collections
Sign up for free to add this paper to one or more collections.